Podospora anserina Genome Project
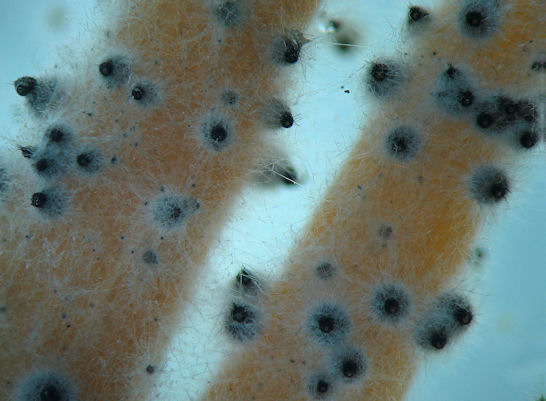 |
Podospora anserina is a filamentous fungus used in several molecular genetics laboratories to study various aspects of mycology and fundamental cell biology (ageing, differentiation and cell death, multicellularity, epigenetics...). This fungus is very helpful as a model system because of the simplicity of its culture, the speed and efficiency of its genetics analysis and the possibility to do molecular biology experiments. |
The complete sequences of the nuclear and mitochondrial genomes of strain S mat+ have been determined in collaboration with CNRS and Genoscope (see history). Microarray generation was funded by ANR.
On this site, sequence data are available for BLAST search and download and microarray data will be posted (see download).
Release
[26 Nov 2013] Release version 6.32 : Assembly of 10x coverage now contains 8 Super contigs. This new release contains a set of 10639 predicted CDS and will be updated periodically.
Funding
The draft sequence was established through fundings from CNRS/Ministère de la recherche, "Séquençage à grande échelle 2002" and Génoscope. Bio-informatic tools were funded by IFR115 Génome. The finishing is funded by IFR115 Génome and Génoscope. Microarrays are funded by ANR (Grant n°ANR-05-Blan-0385).
 |  |  |  |
Contacts
Project coordinator : Philippe Silar
Project management : Robert Debuchy
Website and in silico genome analysis : Olivier Lespinet